In 2020, I was a member of the 2020 UCL iGEM team working on computational modeling, synthetic biology, and website development. The International Genetically Engineered Machine competition (iGEM) is an annual, world wide synthetic biology competition for undergraduates as well as high school students and overgraduates. Our team’s project aimed to tackle two global crises, freshwater scarcity and plastic pollution, by integrating enzymatic polyethylene terephthalate (PET) into a Microbial Desalination Cell (MDC).
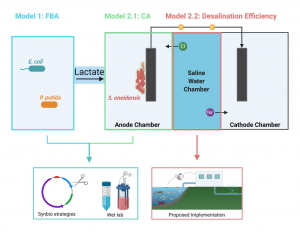
Figure 1. Project overview schematic
As part of the synthetic biology group dedicated to designing constructs, I learned to use Benchling for designing constructs that would allow our bacteria to acquire the desired functionalities to fully degrade PET and feed an exoelectrogenic bacterial strain in the MDC. To study the structures of our plastic degrading enzymes, I received trainings on Pymol and ChimeraX for structural analysis. For structural predictions, Phyre2 and SWISS-MODEL were used.
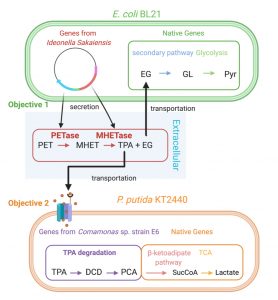
Figure 2. Enzymatic plastic degradation pathway
Moreover, with team support, I performed Flux Balance Analysis (FBA) to simulate and optimize the biomass growth rate as well as product secretion rate of our plastic degrading bacteria. The MATLAB COBRA toolbox was employed for this purpose. The FBA results not only validated the metabolic pathway designed by our synthetic biology group, but also provided information on gene deletions for the optimization of product secretion.
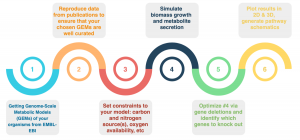
Figure 3. FBA workflow
Finally, to present our achievements to the public and iGEM judges, I led the web development of our team. This stage of work involved graphic design, video editing, and web programming. For producing schematics, I familiarized myself with Biorender. Web development was based on coding in HTML, CSS, and Javascript. Here are some advice on Wiki development:
Figure 4. Advice on Wiki development
For more information our project, please visit our team website. See “Attributions” page for the details of my role.
Acknowledgement: Sincerely appreciate all the support from my teammates and supervisors 🥰.