When you Google the terms “Deep learning and Medicine” or “Machine learning and medicine”, most of the articles you see will probably be about its uses in radiology-based diagnosis, electronic healthcare records, basically the clinical data. Over the past couple of years, there has been increasing ML research on the more cellular level, focusing on genomics and drug interactions at the molecular level. Let’s zoom into the human body and take a look at this!
Biologists have always been aware that cells are highly complex units, but only over the past decade have we actually started to uncover this complexity. This is due to the various improvements in high-throughput techniques which have allowed us to effectively measure and monitor what happens at the cellular level. This area of biological research is broadly called omics and can be broken down into many subtypes. Here are just 3 big ones:
- Genomics – Here, we study the DNA, the molecule that defines everything about us, our hair color, our height, even our risk for disease.
- Proteomics – As you’ll see later, a large part of our body is protein and proteomics is specifically about studying these proteins.
- Metabolomics – This area focuses on how all the important molecules in our body, including our proteins, carbs, and fats, interact with each other in complex chemical reactions, allowing our cells to perform the way they do.
While I’ve defined these areas as different sub-types of omics, in reality, they all overlap to some extent. A vast amount of data is generated in omics, hence computational techniques are regularly used to analyze the data….so regularly used that we have a specific area of research just for this – it’s called bioinformatics (sometimes also synonymous to computational biology). Considering the amount of biological data that exists today, bioinformaticians are in super high demand in medical research.
But while bioinformatics has been around for some time, one of the difficulties for practitioners has been to analyze the data goldmine in an efficient way. The algorithms currently used are computationally expensive and explicitly programmed for conducting data analysis. Instead of this, it would be nice to have a computer learn the patterns in the data. Thus, over the past couple of years, machine learning and deep learning techniques have become popular in bioinformatics. I’d like to show you how ML is being used, but in order to understand the research, we need to first come to grips with the basic biology. Specifically, I’d like to first explore the most important area of omics, genomics. If you’re a biology expert, then you can skip the rest of this post and check out my second post. Otherwise, let’s begin!
Genomics for the beginner
Having gained exposure to genomics during my undergraduate years, I must confess, one thing I’ve struggled with was the large amount of jargon, which goes unexplained and can get frustrating! But I’ll try my best to keep this post jargon-free. So firstly, what is genomics? It’s basically research devoted to understanding the genome – the genome is the cookbook containing recipes to create a functioning living organism, in our case, humans. How is this cookbook organized?
Organization of the Genome
Looking at the structure of any book, the most basic element is the alphabet, right? Combining alphabets together gives words, which together make sentences, then paragraphs, then chapters, and finally the whole book.
The genome’s organization is similar – it’s made up of DNA. DNA is a special molecule which all our cells contain – it defines everything about us, how we look, how our body functions, and whether we are at risk to develop a particular disease.
What does DNA look like? Take a look at the picture of our DNA below. It looks like a spiral ladder right? This structure is famously called the “double helix”.

If we just unwind the double helix structure and zoom in (click on the video below to see this), you will notice that the structure is made up of 2 strands. These strands are made up of 4 types of molecules or 4 nucleotides – adenine, thymine, cytosine, and guanine – they are also known as the bases. The bases are the alphabet of the genomics book and we represent them as the letters “A”, “T”, “C”, and “G”.
So each strand of DNA can be expressed as a sequence of these 4 bases. Now just as an example, here’s a random DNA sequence that I made up:
ATGGACCAGGAAAATGCACTGAACCGTGGCCATAAGGACACATACGAGCAGGATAG
But wait, DNA is made up of 2 strands and I’ve written down just one strand. What is the second strand? Well, from early research by Watson and Crick, we learned that these 4 bases pair up together like two magnets by a phenomenon called "complementary base pairing". The rule is that if one strand has an "A", then the other strand will have a "T" at that position, and if there’s a "C" on one strand, then it will pair with a "G". Keeping this rule in mind, let’s figure out the second DNA strand sequence based on the sequence of the first. The entire DNA sequence will be:
Strand 1 - ATGGACCAGGAAAATGCACTGAACCGTGGCCATAAGGACACATACGAGCAGGATAG
||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Strand 2 - TACCTGGTCCTTTTACGTGACTTGGCACCGGTATTCCTGTGTATGCTCGTCCTATC
Great, we’ve learned the alphabet, now let’s build the book!
Building the book…
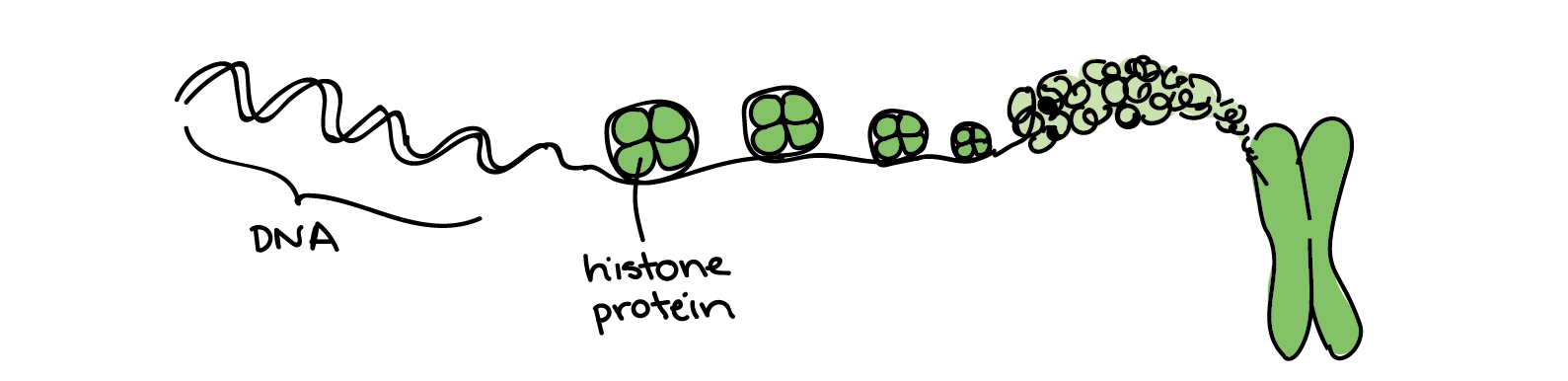
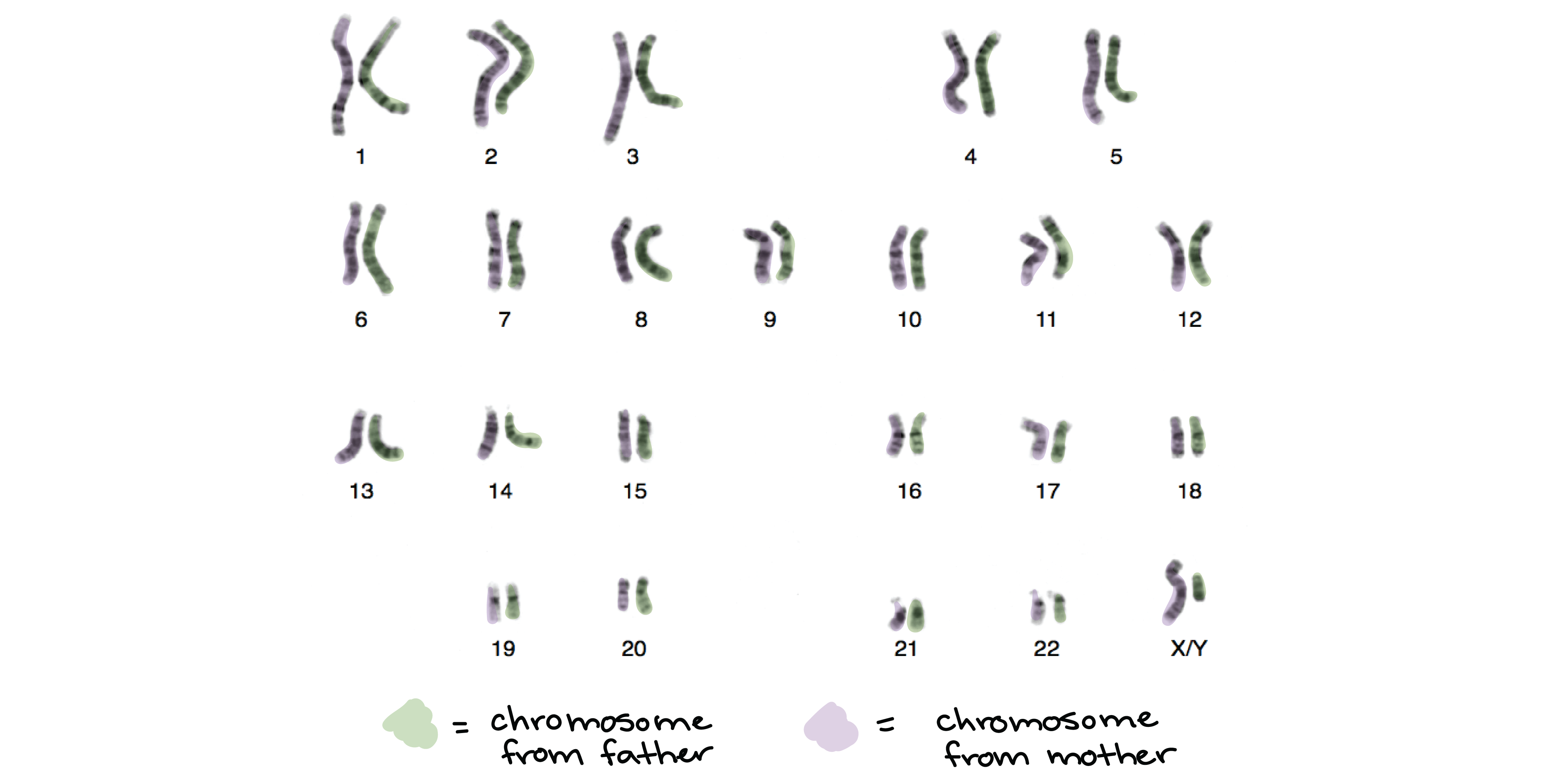
The above DNA sequence I made was only 56 base pairs long. The whole human genome is much much bigger, roughly 3.2 billion base pairs long! That’s a lot of bases, and with a cell being so small, it would be messy to just have all the DNA running like loose threads. So there must be a way to condense this DNA so that it is compact. This is done by basically coiling up the DNA like how you coil your laptop charger when packing it into your backpack. By coiling it further and further, it condenses until it ultimately forms structures called chromosomes, which you may have heard about. Chromosomes are like the chapters in the book. In our genome, we have 23 pairs of chromosomes (23 chapters), of which one pair comes from our mother and the second pair from our father. (If you’re interested, there’s a book by Matt Ridley which actually covers everything about our genome in 23 chapters.)
From DNA to our entire body
So you now know about how the genome is organized! But how exactly does DNA create our physical features? Well, how do you cook a meal from a complex recipe in a cookbook? You read through the recipe, keep your ingredients ready and then physically combine them in the kitchen to make the dish. Putting this in genomics language, the process of “reading” the DNA is called transcription. During transcription, what happens is special machinery in the cell, called RNA polymerase, converts the DNA into another molecule called messenger RNA (mRNA), which is basically a summarized transcript of what has been read.
Finally, this mRNA transcript is passed to another special machinery (called ribosomes) in the cell which does the actual cooking. This process is called translation. Amino acids, which are the basic building blocks of proteins, are combined together like ingredients by the ribosomes to create proteins, the final dish. These proteins will combine to make up components of our cells, the cells combine to make up our tissues, then organs, and finally our entire body! So as you can see, proteins are quite important for us, and in order to make these proteins, DNA is important.
But does all DNA create protein?
If you’ve made it to this point, congratulations because now you covered the fundamentals of genomics! You now know why DNA is called the hereditary molecule, because it creates the proteins that make our organs. But now taking this to the next level, does all DNA make protein? The answer is no. What you’ll find surprising is that in our genome, only 1-2% of it actually codes for protein, while the rest 98-99% doesn’t make any protein or is non-coding!
This was a really important question to ask because it defines what genomics is all about. While reading this post, you may have wondered whether genomics and genetics are the same. They are not! The reason for this distinction is that genetics just focuses on the genes. Genes represent the inherited DNA that creates our physical features. Our physical features are created through the proteins that are coded by the 1-2% of the genome, so by that logic, genetics focuses on just the 1-2% of the genome (which is still huge since we have around 20,000-30,000 genes).
Genomics is broader; it focuses on all the DNA we have, whether it makes a protein or not. Now you may be thinking this 98-99% of DNA that doesn’t make protein is probably useless and junk, which is what scientists thought as well. But over the past decade of research, it has become clear that this non-coding DNA is crucial for helping our cells function. We now know that it controls how our genes switch on/off and how transcription occurs in cells, also called gene expression. So if non-coding DNA is messed up, then protein production can go haywire, eventually leading to life-threatening diseases like cancer. I find this quite funny because it shows how the meta-data contains more information than the data itself. Understanding this better will help us find effective drugs for treating these diseases.
I’m sure you can see now that genomics is really complex, but also super exciting because biologists are trying to uncover the mechanisms of our body, which happen at the molecular level. It is one of the hotspots of biomedical research today, and with so much data being produced, computational approaches like machine learning and bioinformatics are essential to analyzing the data, which is what I’ll discuss more in a future post. Stay tuned!